Reprinted with permission from Quanta Magazine’s Abstractions blog.
In the closing paragraph of On the Origin of Species, Charles Darwin urged readers to “contemplate a tangled bank, clothed with many plants of many kinds, with birds singing on the bushes, with various insects flitting about, and with worms crawling through the damp earth.” Those plants, birds, insects and worms, he continued, all evolved as they did because of the complex web of ecological factors in which they’d been embedded. Had the temperature been hotter, the water more acidic or a certain species of grass absent, a very different “tangled bank” might have evolved instead.
Researchers have typically tried to tease out the evolutionary effects of environmental factors one by one. But the full biodiversity of an environment—the entirety of the tangled bank itself—can also be a crucial influence on how a species evolves.
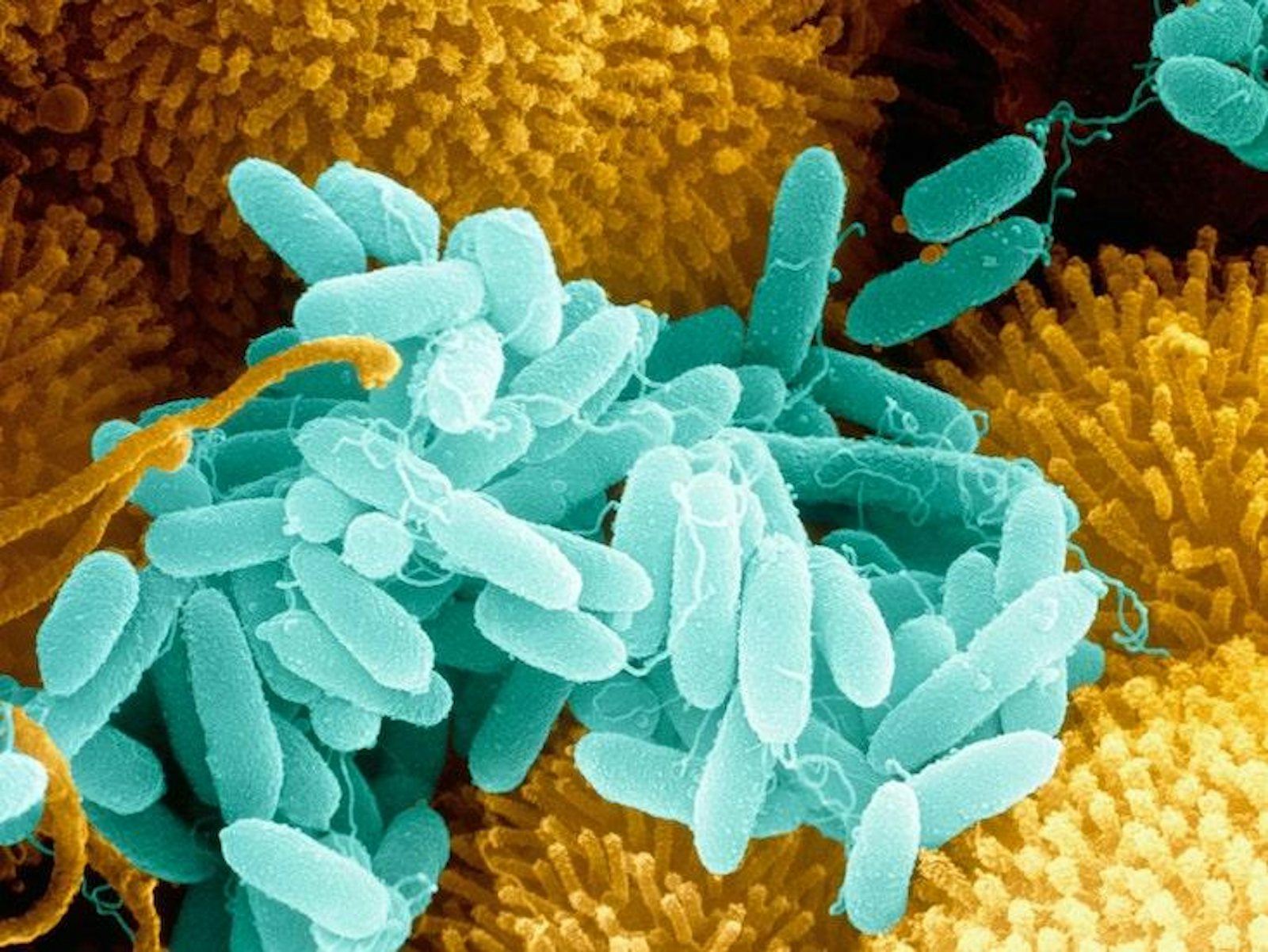
Recently, a paper published in Nature found that when a bacterial species resides in even a very simple ecological community — one that includes just a few other kinds of microbes — it evolves very different defense strategies against a predatory bacteriophage virus than it does when it’s left alone with the phage. The research “is expanding these ideas into the context of the microbiome, where bacteria exist alongside loads of other species,” said Michael Brockhurst, an evolutionary biologist at the University of Sheffield in England who was not involved with the study.
The finding not only elevates the value of biodiversity as an evolutionary factor in its own right, but suggests that some earlier conclusions about the behaviors and capabilities of microorganisms, drawn from laboratory studies of species in isolation, may be seriously incomplete. It also sounds a note of caution about some contemplated strategies for beating drug resistance in bacteria.
Resistance by Any Means Necessary
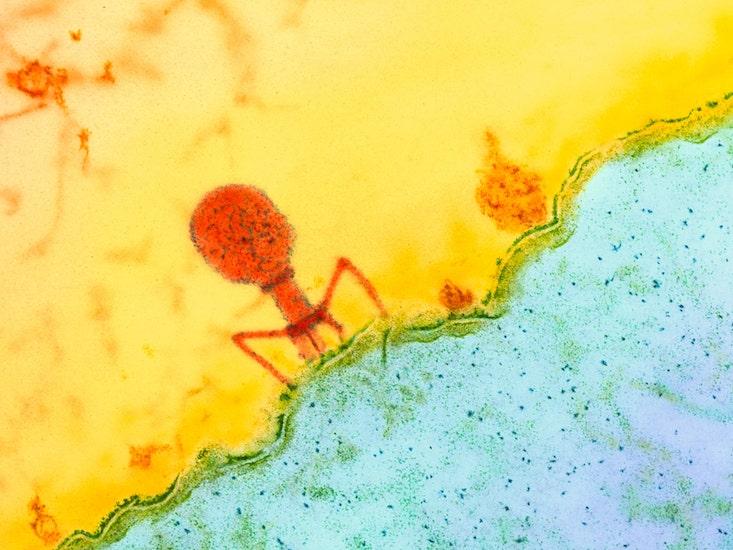
Scientists often turn to bacteria and the bacteriophage viruses that prey on them to learn about coevolution. Both are locked in an eternal, rapid-fire evolutionary conflict that can be controlled and measured in the laboratory.
Eons ago, bacteria (or their archaea cousins) evolved a clever system for thwarting viruses: the genomic feature called CRISPR, which has become famous in the past decade as a tool of biotechnology. CRISPR acts like an adaptive immune system; it enables bacteria that have been exposed to a virus to pass on a genetic “memory” of that infection to their descendants, which can then mount better defenses against a repeat infection. It’s a system that works so well that an estimated half of all bacterial species use CRISPR.
What puzzles many microbiologists, though, is why some bacteria use CRISPR and others don’t. CRISPR-equipped bacteria are scattered throughout the bacterial kingdom almost randomly, and even the bacteria with CRISPR in their genomes don’t consistently rely on it.
Researchers have uncovered dozens of other systems that bacteria use to rebuff phage invasions. But in laboratory studies, bacteria primarily develop what’s known as surface-based phage resistance. Mutations change receptor molecules on the surface of the bacterial cell, so that the phage can no longer recognize and invade it.
The strategy is akin to shutting a door and throwing away the key: It offers the bacteria complete safety from infection by the virus. But that protection comes at a significant price, because it also disrupts whatever nutrient uptake, waste disposal, communication task or other cellular function the receptor would have been providing—taking a constant toll on a cell’s fitness.
In contrast, CRISPR only drags on a cell’s resources when it’s active, during a viral infection. Even so, CRISPR represents a riskier gambit: It doesn’t start to work until phages have already entered the cell, meaning that there’s a chance the viruses could overcome it. And CRISPR doesn’t just attack viral DNA; it can also prevent bacteria from taking up beneficial genes from other microbes, like those that confer antibiotic resistance.
What factors affect the trade-offs in costs and fitness? For the past six years, Edze Westra, an evolutionary ecologist at the University of Exeter in England, has led a team pursuing the answer to that question. In 2015, they discovered that nutrient availability and phage density affected whether Pseudomonas bacteria relied on surface-based or CRISPR-based resistance. In environments poor in resources, receptor modifications were more burdensome, so CRISPR became a better bargain. When resources were plentiful, bacteria grew more densely and phage epidemics became more frequent. Bacteria then faced greater selective pressure to close themselves off from infection entirely, and so they shut down receptors to gain surface-based resistance.
This explained why surface-based resistance was so common in laboratory cultures. Growing in a test tube rich in nutrients, “these bacteria are on a holiday,” Westra said. “They are having a terrific time.”
Still, these rules weren’t cut and dried. Plenty of bacteria in natural high-nutrient environments use CRISPR, and plenty of bacteria in natural low-nutrient environments don’t. “It’s all over the place,” Westra said. “That told us that we were probably still missing something.”
How Biodiversity Reshapes the Battle
Then one of Westra’s graduate students, Ellinor Opsal, proposed another potential factor: the diversity of the biological communities in which bacteria live. This factor is harder to study, but scientists had previously observed that it could affect phage immunity in bacteria. For example, in 2005, James Bull, a biologist at the University of Texas, Austin, and William Harcombe, his graduate student at the time (now at the University of Minnesota), found that E. coli bacteria didn’t evolve immunity to a phage when a second bacterial species was present. Similarly, Britt Koskella, an evolutionary biologist at the University of California, Berkeley, and one of her graduate students, Catherine Hernandez, reported last year that phage resistance failed to arise in Pseudomonas bacteria living on their actual host (a plant), though they always gained immunity in a test tube. Could the diversity of the surroundings influence not just whether or not resistance to phages evolved, but the nature of that resistance?
To find out, Westra’s team performed a new set of experiments: Instead of altering the nutrient conditions for Pseudomonas bacteria growing with phages, they added three other bacterial species — species that competed against Pseudomonas for resources but weren’t targeted by the phage.
Left to themselves, Pseudomonas would normally develop surface-based mutations. But in the company of rivals, they were far more likely to turn to CRISPR. Further investigation showed that the more complex community dynamics had shifted the fitness costs: The bacteria could no longer afford to inactivate receptors because they not only had to survive the phage, but also had to outcompete the bacteria around them.
These results from Westra’s group dovetail with earlier findings that phages can produce greater diversity in bacterial communities. “Now, that diversity is actually feeding back to the phage side of things” by affecting phage resistance, Koskella said. “It’s neat to see that coming full circle.” By understanding that kind of feedback loop, she added, “we can start to ask more general questions about the impacts that phages have in a community context.”
For one, the bacteria’s shift toward a CRISPR-based phage response had another, broader effect. When Westra’s group grew Pseudomonas in moth larvae hosts, they found that the bacteria with surface-based resistance were less virulent, killing the larvae much more slowly than the bacteria with active CRISPR systems did.
These results have direct implications for phage therapy, an approach to fighting bacterial infections that is gaining interest among researchers. Scientists like Paul Turner, an ecologist and evolutionary biologist at Yale University, are seeking out phages that would induce receptor mutations in targeted bacteria to make them less virulent or more susceptible to antibiotics. But if the evolution of CRISPR-based resistance is an option for the bacteria, that strategy “might not always work in a more complex community,” Turner said. (He acknowledged that this hasn’t yet surfaced as a problem in phage therapy experiments).
Westra’s team, along with other groups, is now studying these effects in other bacterial systems and environments, and for other types of immunity. They’re also exploring how different types of microbial diversity affect the evolution of phage resistance in bacteria. Meanwhile, Harcombe and his colleagues are studying how evolution proceeds in communities where bacterial species cooperate and rely on each other for survival instead of competing, such as the microbiome in the human gut.
Others are looking at evolutionary traits beyond phage resistance. In a paper published in August, a team of researchers found that greater biodiversity prevented the spread of certain antibiotic resistance genes.
Perhaps most important, these findings give scientists insights into the level of complexity they might need for their experiments going forward. “A lot of the stuff that we think we know about bacteria and phage and how they interact comes from these really simple test tube experiments,” Westra said. “And these results just don’t hold if we start to introduce some real-life ecological complexity. We have to take the environmental realism into account”—even if it is a much more difficult endeavor.
“That’s going to be really critical,” Harcombe said, “as we try to manage microbiomes and as we try to manage ecosystems in a changing world.”
Jordana Cepelewicz is a staff writer at Quanta Magazine who covers biology.