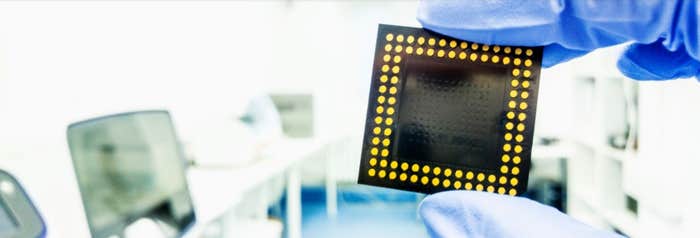
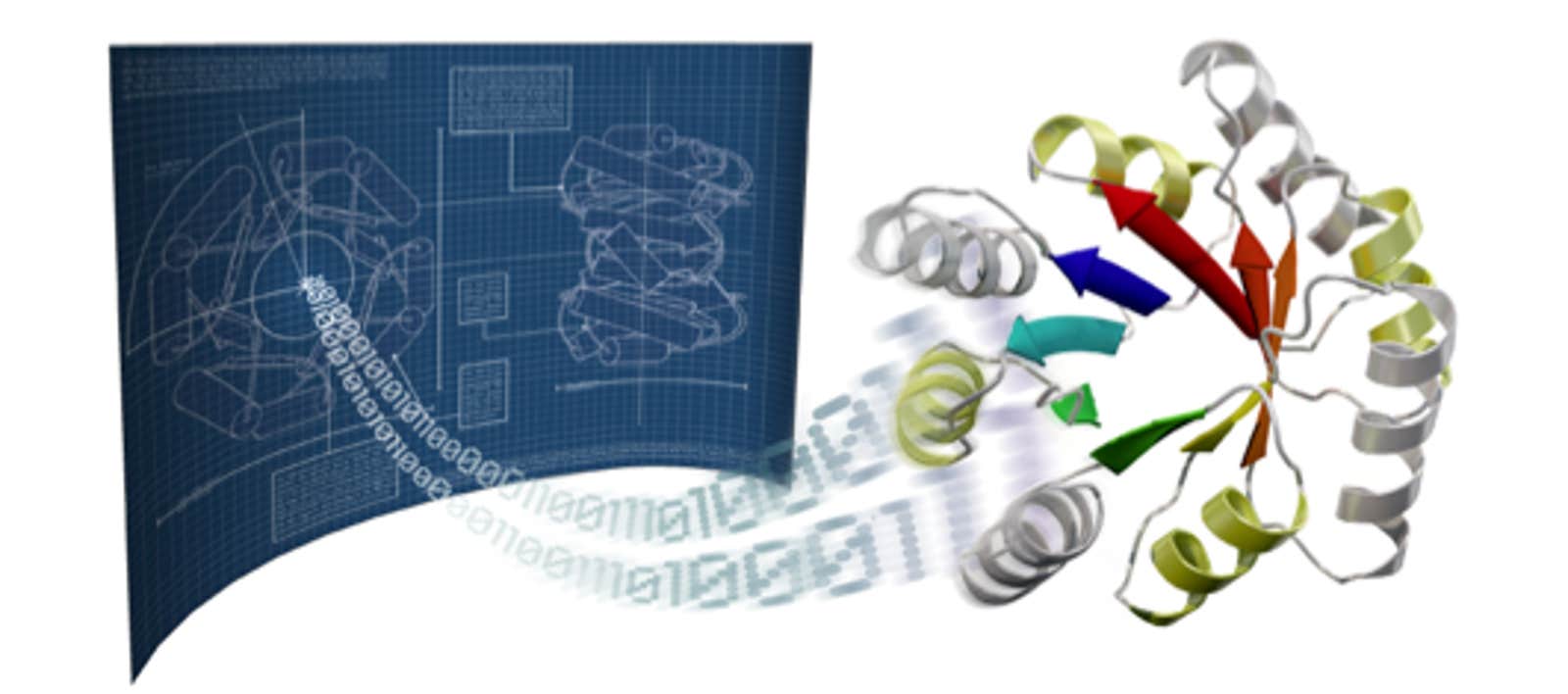
Proteins are the workhorses of all living creatures, fulfilling the instructions of DNA. They occur in a wide variety of complex structures and carry out all the important functions in our body and in all living organisms—digesting food, building tissue, transporting oxygen through the bloodstream, dividing cells, firing neurons, and powering muscles. Remarkably, this versatility comes from different combinations, or sequences, of just 20 amino acid molecules. How these linear sequences fold up into complex structures is just now beginning to be well understood (see box).
Even more remarkably, nature seems to have made use of only a tiny fraction of the potential protein structures available—and there are many. Therein lies an amazing set of opportunities to design novel proteins with unique structures: synthetic proteins that do not occur in nature, but are made from the same set of naturally-occurring amino acids. These synthetic proteins can be “manufactured” by harnessing the genetic machinery of living things, such as in bacteria given appropriate DNA that specify the desired amino acid sequence. The ability to create and explore such synthetic proteins with atomic level accuracy—which we have demonstrated—has the potential to unlock new areas of basic research and to create practical applications in a wide range of fields.
The design process starts by envisioning a novel structure to solve a particular problem or accomplish a specific function, and then works backwards to identify possible amino acid sequences that can fold up to this structure. The Rosetta protein modelling and design software identifies the most likely candidates—those that fold to the lowest energy state for the desired structure. Those sequences then move from the computer to the lab, where the synthetic protein is created and tested—preferably in partnership with other research teams that bring domain expertise for the type of protein being created.
At present no other advanced technology can beat the remarkable precision with which proteins carry out their unique and beautiful functions. The methods of protein design expand the reach of protein technology, because the possibilities to create new synthetic proteins are essentially unlimited. We illustrate that claim with some of the new proteins we have already developed using this design process, and with examples of the fundamental research challenges and areas of practical application that they exemplify:
Catalysts for clean energy and medicine. Protein enzymes are the most efficient catalysts known, far more so than any synthesized by inorganic chemists. Part of that efficiency comes from their ability to accurately position key parts of the enzyme in relation to reacting molecules, providing an environment that accelerates a reaction or lowers the energy needed for it to occur. Exactly how this occurs remains a fundamental problem which more experience with synthetic proteins may help to resolve.
Already we have produced synthetic enzymes that catalyze potentially useful new metabolic pathways. These include: reactions that take carbon dioxide from the atmosphere and convert it into organic molecules, such as fuels, more efficiently than any inorganic catalyst, potentially enabling a carbon-neutral source of fuels; and reactions that address unsolved medical problems, including a potential oral therapeutic drug for patients with celiac disease that breaks down gluten in the stomach and other synthetic proteins to neutralize toxic amyloids found in Alzheimer’s disease.
We have also begun to understand how to design, de novo, scaffolds that are the basis for entire superfamilies of known enzymes (Fig. 1) and other proteins known to bind the smaller molecules involved in basic biochemistry. This has opened the door for potential methods to degrade pollutants or toxins that threaten food safety.
New super-strong materials. A potentially very useful new class of materials is that formed by hybrids of organic and inorganic matter. One naturally occurring example is abalone shell, which is made up of a combination of calcium carbonate bonded with proteins that results in a uniquely tough material. Apparently, other proteins involved in the process of forming the shell change the way in which the inorganic material precipitates onto the binding protein and also help organize the overall structure of the material. Synthetic proteins could potentially duplicate this process and expand this class of materials. Another class of materials are analogous to spider silk—organic materials that are both very strong and yet biodegradable—for which synthetic proteins might be uniquely suited, although how these are formed is not yet understood. We have also made synthetic proteins that create an interlocking pattern to form a surface only one molecule thick, which suggest possibilities for new anti-corrosion films or novel organic solar cells.
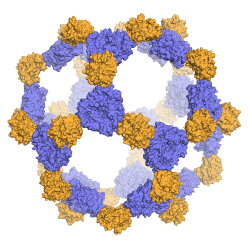
Targeted therapeutic delivery. Self-assembling protein materials make a wide variety of containers or external barriers for living things, from protein shells for viruses to the exterior wall of virtually all living cells. We have developed a way to design and build similar containers: very small cage-like structures—protein nanoparticles—that self-assemble from one or two synthetic protein building blocks (Fig. 2). We do this extremely precisely, with control at the atomic level. Current work focuses on building these protein nanoparticles to carry a desired cargo—a drug or other therapeutic—inside the cage, while also incorporating other proteins of interest on their surface. The surface protein is chosen to bind to a similar protein on target cells.
These self-assembling particles are a completely new way of delivering drugs to cells in a targeted fashion, avoiding harmful effects elsewhere in the body. Other nanoparticles might be designed to penetrate the blood-brain barrier, in order to deliver drugs or other therapies for brain diseases. We have also generated methods to design proteins that disrupt protein-protein interactions and proteins that bind to small molecules for use in biosensing applications, such as identifying pathogens. More fundamentally, synthetic proteins may well provide the tools that enable improved targeting of drugs and other therapies, as well as an improved ability to bond therapeutic packages tightly to a target cell wall.

Novel vaccines for viral diseases. In addition to drug delivery, self-assembling protein nanoparticles are a promising foundation for the design of vaccines. By displaying stabilized versions of viral proteins on the surfaces of designed nanoparticles, we hope to elicit strong and specific immune responses in cells to neutralize viruses like HIV and influenza. We are currently investigating the potential of these nanoparticles as vaccines against a number of viruses. The thermal stability of these designer vaccines should help eliminate the need for complicated cold chain storage systems, broadening global access to life saving vaccines and supporting goals for eradication of viral diseases. The ability to shape these designed vaccines with atomic level accuracy also enables a systematic study of how immune systems recognize and defend against pathogens. In turn, the findings will support development of tolerizing vaccines, which could train the immune system to stop attacking host tissues in autoimmune disease or over-reacting to allergens in asthma.
New peptide medicines. Most approved drugs are either bulky proteins or small molecules. Naturally occurring peptides (amino acid compounds) that are constrained or stabilized so that they precisely complement their biological target are intermediate in size, and are among the most potent pharmacological compounds known. In effect, they have the advantages of both proteins and small molecule drugs. The antibiotic cyclosporine is a familiar example. Unfortunately such peptides are few in number.
We have recently demonstrated a new computational design method that can generate two broad classes of peptides that have exceptional stability against heat or chemical degradation. These include peptides that can be genetically encoded (and can be produced by bacteria) as well as some that include amino acids that do not occur in nature. Such peptides are, in effect, scaffolds or design templates for creating whole new classes of peptide medicines.
In addition, we have developed general methods for designing small and stable proteins that bind strongly to pathogenic proteins. One such designed protein binds the viral glycoprotein hemagglutinin, which is responsible for influenza entry into cells. These designed proteins protect infected mice in both a prophylactic and therapeutic manner and therefore are potentially very powerful anti-flu medicines. Similar methods are being applied to design therapeutic proteins against the Ebola virus and other targets that are relevant in cancer or autoimmune diseases. More fundamentally, synthetic proteins may be useful as test probes in working out the detailed molecular chemistry of the immune system.
Protein logic systems. The brain is a very energy-efficient logic system based entirely on proteins. Might it be possible to build a logic system—a computer—from synthetic proteins that would self-assemble and be both cheaper and more efficient than silicon logic systems? Naturally occurring protein switches are well studied, but building synthetic switches remains an unsolved challenge. Quite apart from bio-technology applications, understanding protein logic systems may have more fundamental results, such as clarifying how our brains make decisions or initiate processes.
The opportunities for the design of synthetic proteins are endless, with new research frontiers and a huge variety of practical applications to be explored. In effect, we have an emerging ability to design new molecules to solve specific problems—just as modern technology does outside the realm of biology. This could not be a more exciting time for protein design.
Predicting Protein Structure
If we were unable to predict the structure that results from a given sequence of amino acids, synthetic protein design would be an almost impossible task. There are 20 naturally-occurring amino acids, which can be linked in any order and can fold into an astronomical number of potential structures. Fortunately the structure prediction problem is now well on the way toward being solved by the Rosetta protein modeling software.
The Rosetta tool evaluates possible structures, calculates their energy states, and identifies the lowest energy structure—usually, the one that occurs in a living organism. For smaller proteins, Rosetta predictions are already reasonably accurate. The power and accuracy of the Rosetta algorithms are steadily improving thanks to the work of a cooperative global network of several hundred protein scientists. New discoveries—such as identifying amino acid pairs that co-evolve in living systems and thus are likely to be co-located in protein structures—are also helping to improve prediction accuracy.
Our research team has already revealed the structures for more than a thousand protein families, and we expect to be able to predict the structure for nearly any protein within a few years. This is an important achievement with direct significance for basic biology and biomedical science, since understanding structure leads to understanding the function of the myriad proteins found in the human body and in all living things. Moreover, predicting protein structure is also the critical enabling tool for designing novel, “synthetic” proteins that do not occur in nature.
How to Create Synthetic Proteins that Solve Important Problems

Now that it is possible to design a variety of new proteins from scratch, it is imperative to identify the most pressing problems that need to be solved, and focus on designing the types of proteins that are needed to address these problems. Protein design researchers need to collaborate with experts in a wide variety of fields to take our work from initial protein design to the next stages of development. As the examples above suggest, those partners should include experts in industrial scale catalysis, fundamental materials science and materials processing, biomedical therapeutics and diagnostics, immunology and vaccine design, and both neural systems and computer logic. The partnerships should be sustained over multiple years in order to prioritize the most important problems and test successive potential solutions.
A funding level of $100M over five years would propel protein design to the forefront of biomedical research, supporting multiple and parallel collaborations with experts worldwide to arrive at breakthroughs in medicine, energy, and technology, while also furthering a basic understanding of biological processes. Current funding is unable to meet the demands of this rapidly growing field and does not allow for the design and production of new proteins at an appropriate scale for testing and ultimately production, distribution, and implementation. Private philanthropy could overcome this deficit and allow us to jump ahead to the next generation of proteins—and thus to use the full capacity of the amino acid legacy that evolution has provided us.